A question that has come up a lot on the blog is what endocast fossils can tell us about the brains and behaviors of long-extinct animals. This question is especially salient for Homo naledi, an unexpected human cousin that lived in South Africa at the same time as the earliest modern-like humans around 300,000 years ago. Brain size in H. naledi ranged from 460–610 ml, similar to Australopithecus and the earliest Homo over 1.5 million years ago and less than half the size of other, contemporaneous fossil humans. Despite its small brain size, the frontal lobe of H. naledi seems to have been more similar to humans than australopiths, including in the area associated with speech.
To learn more about the brain of Homo naledi, I teamed up with Shawn Hurst and John Hawks to virtually reconstruct the endocast from the most complete skull and skeleton of the species, nicknamed Neo (which means “gift” in the SeSotho language). We have a paper about it coming out soon in the journal Brain Structure and Function, where we show that Neo’s endocast shape is fairly distinct among Pleistocene hominins. We go on to discuss some implications and limitations of our results for understanding the brain and behavior of H. naledi and other hominins. I’ll write more about it when the paper is actually in press, but in the meantime you can get a sneak preview by checking the data and running the analyses for yourself.
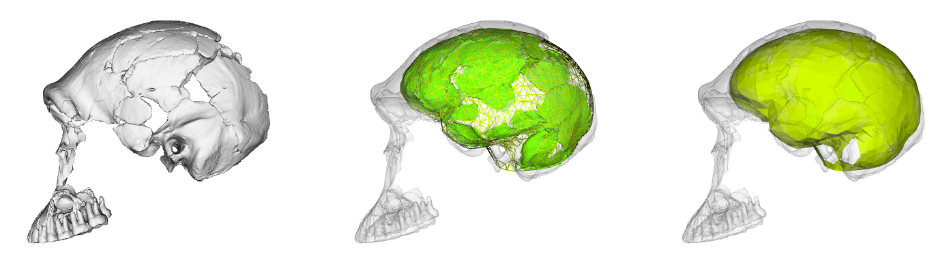
I published the landmark data and R code from our study in the open access repository Zenodo (here). I built off the endocast landmark data that Simon Neubauer and colleagues (2018) made available from their study of the KNM-ER 42700 fossil. I created a landmark template from that dataset and applied it to Neo and Australopithecus africanus cranium Sts 5, and then used geometric morphometric methods to reconstruct the missing regions of Neo and compare it to the other hominins. The accompanying R code walks you through inputting these data, estimating missing landmark positions, and comparing endocast shapes of humans and fossil hominins.

Hopefully these data and code will let others build off our results, add more fossils to the mix, and generate more insights about how the human brain has changed over the past several millions of years.