UPDATE: sadly, the KUPRI Digital Morphology Museum website, which was the basis for this activity, is no longer online 😦 If I can find a comparable resource I will be sure to update this post.
The focus of my human evolution class the past few weeks has been uncovering the earliest human ancestors. The main adaptation distinguishing our first forebears from other animals is walking on two legs (“bipedalism”), so researchers try to identify features reflective of bipedalism in fossils over 4 million years ago. But this isn’t so easy – not all fossils will tell us how an animal walked around, and even with the right bones, it’s not always clear what the earliest bipeds “should” look like. Take the case of Ardipithecus kadabba: there are a handful of seemingly nondescript fossils (below) from a number of Ethiopian sites dating 5.2-5.8 million years ago. Can we really tell if this species was bipedal? LET YOUR STUDENTS DO SCIENCE TO DECIDE FOR THEMSELVES!

Can you spot a biped? Ardipithecus kadabba fossils. Scale bar is 1 cm (Fig. 1 from Haile-Selassie, 2001).
In the jumble of fragments pictured above, the key bone possibly revealing a bipedal animal is in square b, a toe bone shown in several views. This is a fourth proximal pedal phalanx (PPP4), where a wedding ring would sit if people put rings on their feet instead of their hands. Ew. Anyway, here’s closer view, from the Ar. kadabba monograph (Haile-Selassie and WoldeGabriel, 2009):

Top: AME-VP 1/71, the sciencey name of the toe bone in question. The proximal end (toward the foot) is to the left and the distal end (toward the tip of the toe), is to the right. Bottom: Lookit how tiny it is! Modified from plates 7.8 and 7.21 in the monograph.
Although absolutely small, the kadabba PPP4 is relatively long, narrow and curved, like an ape’s and unlike a human’s. Haile-Selassie and colleagues (monograph chapter) compared this fossil with chimpanzee and the bipedal Australopithecus afarensis‘s PPP4s (below I have scaled them to the same length). As you can see, kadabba and the chimp are fairly narrow compared to the stout afarensis toe, although they are all curved:

Comparison of PPP4s, scaled to same length. Left to right, and top to bottom: Chimp, Ar. kadabba and Au. afarensis. The left box is a bottom view (proximal at the bottom) and the right box from the side (proximal to the left). Modified from plates 7.24-7.25 in the kadabba monograph.
These gross shape comparisons don’t make kadabba‘s toe look that like of a bipedal animal. However, one thing we can’t see in these views – and that you can have your students examine in virtual lab – is the orientation of the proximal joint surface. In humans (Griffin and Richmond, 2010) and the later Ardipithecus ramidus and australopithecines (Latimer and Lovejoy 1990; Lovejoy et al., 2009; Haile-Selassie et al., 2012), this joint surface angles upward, a result of the great force this joint experiences as it hyper-dorsiflexes during walking. Apes and other animals are not bipedal, and they do not dorsiflex their toes to the degree that we do, so this joint does not usually angle upward as much.
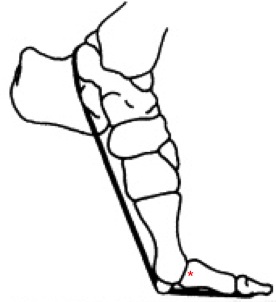
Schematic of a hyper-dorseflexed proximal toe joint (indicated by red star).
Now here’s that joint in the kadabba PPP4. The kadabba monograph has a nice midsagittal CT scan revealing this joint’s orientation, the angle of which I have measured using the free image analysis software ImageJ. kadabba‘s angle of ‘dorsal canting’ is 102.2 degrees. A mild fever.

Measuring the dorsal canting (the angle theta) of the AME-VP-1/71 proximal joint surface, using ImageJ.
This measurement is at the low end of the human range (averaging around 110 degrees for the 2nd, not 4th, digit), and above all but just a few apes analyzed by Griffin and Richmond (2010). Now, these authors looked at PPP2s, but PPP4s would be more appropriate for comparison with kadabba. LUCKY YOU – your students can collect these data, using CT scans from the KUPRI digital morphology museum. This collection has dozens of apes and monkeys (and even some other mammals), presumably none of which were habitually bipedal, and which should have relatively low angles of canting. (many of these specimens are from zoos, however, so their activity patterns and anatomies may not be the same as wild animals’; lookit the gorilla below) Your students can isolate and section proximal pedal phalanges as I have below, and measure the angle of canting with ImageJ. SCIENCE!

Easy image acquisition on the KUPRI database (this is a gorilla, with a pretty messed up fourth digit beyond the distal PPP). Have your students save the sectioned image as above, then measure the angle theta.
This activity is simple way for your students to set up a hypothesis, collect quality data and analyze them – essentially for free!
Some light weekend reading
Griffin and Richmond (2010). Joint orientation and function in great ape and human proximal pedal phalanges. American Journal of Physical Anthropology 141: 116-123.
Haile-Selassie (2001). Late Miocene hominids from the Middle Awash, Ethiopia. Nature 412: 178-181.
Haile-Selassie and WoldeGabriel, eds. (2009) Ardipithecus kadabba: Late Miocene Evidence from the Middle Awash, Ethiopia. Chapter 7.
Haile-Selassie et al. (2012). A new hominin foot from Ethiopia shows multiple Pliocene bipedal adaptations. Nature 483: 565-570.
Latimer and Lovejoy (1990). Metatarsophalangeal joints of Australopithecus afarensis. American Journal of Physical Anthropology 83: 13-23.
Lovejoy et al. (2009). Combining prehension and propulsion: The foot of Ardipithecus ramidus. Science 326: 72e1-72e8