The annual meetings of the American Association of Physical Anthropologists were going on all last week, and I gave my first talk before the Association (co-authored with Jeremy DeSilva). The talk focused on using resampling methods and the abysmal human fossil record to assess whether human-like brain size growth rates were present in our >1 mya ancestor Homo erectus. This is something I’ve actually been sitting on for a while, and wanted to wait til after the talk to post for all to see. I haven’t written this up yet for publication, but before then I’d like to briefly share the results here.
Background: Humans’ large brains are critical for giving us our unique capabilities such as language and culture. We achieve these large (both absolutely, and relative to our body size) brains by having really high brain growth rates across several years; most notable are exceptionally high, “fetal-like” rates during the first 1-2 years of life. Thus, rapid brain growth shortly after birth is a key aspect of human uniqueness – but how ancient is this strategy?
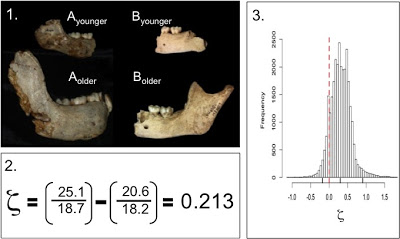
Materials: We can plot brain size at birth in humans and chimpanzees (our closest living relatives) to visualize what makes humans stand out (Figure 1).
 |
| Figure 1. Brain size (volume) at given ages. Humans=black, chimpanzees=red. Ranges of brain size at birth, and the chronological age of the Mojokerto fossil, in blue. |
Human data come from Cogueugniot and Hublin (2012), and chimpanzees from Herndon et al. (1999) and Neubauer et al. (2012). The earliest fossil evidence able to address this question comes from Homo erectus. Because of the tight relationship between newborn and adult brain size (DeSilva and Lesnik 2008), we can use adult Homo erectus brain volumes (n=10, mean = 916.5 cm^3) to predict that of the species’ newborns: mean = 288.9 cm^3, sd = 17.1). An almost-recent analysis of the Mojokerto Homo erectus infant calvaria suggests a size of 663 cm^3 and an age of 0.5-1.25 years (Coqueugniot et al. 2004; this study actually suggests an oldest age of 1.5 years, but the chimpanzee sample here requires us to limit the study to no more than 1.25 years). Because we have a H. erectus fossil less than 2 years of age, and we can estimate brain size at birth, we can indirectly assess early brain growth in this species.
Methods: Resampling statistics allow inferences about brain growth rates in this extinct species, incorporating the uncertainty in both brain size at birth, and in the chronological age of the Mojokerto fossil. We thus ask of each species, what growth rates are necessary to grow one of the newborn brain sizes to any infant between 0.5-1.25 years? And from there, we compare these resampled growth rates (or rather, ‘pseudo-velocities’) between species – is H. erectus more similar to modern humans or chimpanzees? There are 294 unique newborn-infant comparisons for humans and 240 for the chimpanzee sample. We therefore compare these empirical newborn-infant pairs from extant species to 7500 resampled H. erectus pairs, randomly selecting a newborn H. erectus size based on the parameters above, and randomly selecting an age from 0.5-1.25 years for the Mojokerto specimen. This procedure is used to compare both absolute size change (the difference between an infant and a newborn size, in cm^3/year), and and proportional size change (infant/newborn size).
Results: Humans’ high early brain growth rates after birth are reflected in the ‘pseudovelocity curve’ (Figure 2). Chimps have a similar pattern of faster rates earlier on, but these are ultimately lower than humans’. Using the Mojokerto infant’s brain size (and it’s probable ages) and the likely range of H. erectus neonatal brain sizes (mean = 288, sd = 17), it is fairly clear that H. erectus achieved its infant brain size with high, human-like rates in brain volume increase.
 |
| Figure 2. Brain size growth rates (‘pseudo-velocity’) at given ages. Humans=black, chimpanzees=red, and Homo erectus,=blue. |
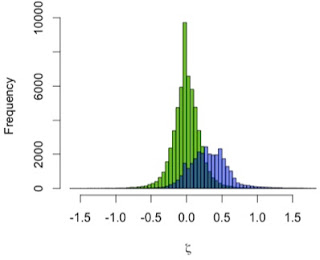
However, if we look at proportional size change, the factor by which brain size increases from birth to a given age, we see a great deal of overlap both between age groups within a species, and between different species. Cross-sectional data create a great deal of overlap in implied proportional size change between ages within a species; it is easier to consider proportional size change between taxa, conflating ages, then (Figure 3). Humans show a massive amount of variation in potential growth rates from birth to 0.5-1.25 years, and chimpanzees also show a great deal of variation, albeit generally lower than in the human sample. Relative growth rates in Homo erectus are intermediate between the two extant species.
 |
| Figure 3. Proportional brain size increase (infant/newborn size). |
Significance: Brain size growth shortly after birth is critical for humans’ adaptative strategy: growing a large brain requires a lot of energy and parental (especially maternal) investment (Leigh 2004). Plus, in humans this rapid increase may correspond with the creation of innumerable white-matter connections between regions of the brain (Sakai et al. 2012), important for cognition or intelligence. The H. erectus fossil record (1 infant and 10 adults) provides a limited view into this developmental period. However, comparative data on extant animals (e.g. brain sizes from birth to adulthood), coupled with resampling statistics, allow inferences to be made about brain growth rates in H. erectus over 1 million years ago.
Assuming the Mojokerto H. erectus infant is accurately aged (Coqueugniot et al. 2004), and that Homo erectus followed the same neonatal-adult scaling relationship as other apes and monkeys (DeSilva and Lesnik 2008), it is likely that H. erectus had human-like rates of absolute brain size growth. Thus, the energetic and parental requirements to raise such brainy babies, seen in modern humans, may have been present in Homo erectus some 1.5 million years ago or so. This may also imply rapid white-matter proliferation (i.e. neural connections) in this species, suggesting an intellectually (i.e. socially or linguistically) stimulating infancy and childhood in this species. At the same time, relative brain size growth appears to scale with overall brain size: larger brains require proportionally higher growth rates. This is in line with studies suggesting that in many ways, the human brain is a scaled-up version of other primates’ (e.g. Herculano-Houzel 2012).

This study was made possible with published data, and the free statistical programming language R.
Contact me if you want the R code used for this analysis, I’m glad to share it!
References
Coqueugniot H, Hublin JJ, Veillon F, Houët F, & Jacob T (2004). Early brain growth in Homo erectus and implications for cognitive ability. Nature, 431 (7006), 299-302 PMID: 15372030
Coqueugniot H, & Hublin JJ (2012). Age-related changes of digital endocranial volume during human ontogeny: results from an osteological reference collection. American journal of physical anthropology, 147 (2), 312-8 PMID: 22190338
DeSilva JM, & Lesnik JJ (2008). Brain size at birth throughout human evolution: a new method for estimating neonatal brain size in hominins. Journal of human evolution, 55 (6), 1064-74 PMID: 18789811
Herculano-Houzel S (2012). The remarkable, yet not extraordinary, human brain as a scaled-up primate brain and its associated cost. Proceedings of the National Academy of Sciences of the United States of America, 109 Suppl 1, 10661-8 PMID: 22723358
Herndon JG, Tigges J, Anderson DC, Klumpp SA, & McClure HM (1999). Brain weight throughout the life span of the chimpanzee. The Journal of comparative neurology, 409 (4), 567-72 PMID: 10376740
Leigh SR (2004). Brain growth, life history, and cognition in primate and human evolution. American journal of primatology, 62 (3), 139-64 PMID: 15027089
Neubauer, S., Gunz, P., Schwarz, U., Hublin, J., & Boesch, C. (2012). Brief communication: Endocranial volumes in an ontogenetic sample of chimpanzees from the taï forest national park, ivory coast American Journal of Physical Anthropology, 147 (2), 319-325 DOI: 10.1002/ajpa.21641
Sakai T, Matsui M, Mikami A, Malkova L, Hamada Y, Tomonaga M, Suzuki J, Tanaka M, Miyabe-Nishiwaki T, Makishima H, Nakatsukasa M, & Matsuzawa T (2012). Developmental patterns of chimpanzee cerebral tissues provide important clues for understanding the remarkable enlargement of the human brain. Proceedings. Biological sciences / The Royal Society, 280 (1753) PMID: 23256194

![]() An important question, and follow-up to Pietak et al’s study, is whether human limb proportions can be described by such log-periodic functions throughout ontogeny, and if so how these change. Plus, it’s also not clear to what extent human proportions might happen to be describable by log periodic functions, simply because each segment is shorter than the one preceding it proximally. In short, this study raises really interesting and pursuable questions about how and why animal limbs grow to the size and proportions that they do.
An important question, and follow-up to Pietak et al’s study, is whether human limb proportions can be described by such log-periodic functions throughout ontogeny, and if so how these change. Plus, it’s also not clear to what extent human proportions might happen to be describable by log periodic functions, simply because each segment is shorter than the one preceding it proximally. In short, this study raises really interesting and pursuable questions about how and why animal limbs grow to the size and proportions that they do.